How Proteins Find and Bind Each Other
Scientists from Freie Universität Berlin and University Pompeu Fabra Barcelona reveal the dynamical mechanism of protein association in atomic detail
№ 167/2017 from Jun 22, 2017
Scientists from Freie Universität Berlin and University Pompeu Fabra (UPF) Barcelona, have for the first time, simulated the association and dissociation of protein molecules in atomic detail, and validated their results with experimental data. “The simulations revealed many previously unknown details on how proteins find and bind each other. But the most valuable result of our study is the demonstration that studying protein-protein association in atomic detail is possible,” explains Dr. Nuria Plattner, a postdoctoral researcher at Freie Unievrsität Berlin and lead author of the study. This achievement is useful not just for biology textbooks, but it also opens the door to understanding the details of viral infections, the inner workings of the immune system, and many other problems with biomedical or biotechnological relevance. The results were published in the highly reputable scientific journal Nature Chemistry.
“Proteins and other molecules move around in our cells, and they can associate to and dissociate from each other,” explains Prof. Frank Noé, a computational scientist at Freie Unversität and mathematician at Reseach Center Matheon who led the research project at Freie Universität Berlin. Human muscles, for example, consist of proteins that have associated to each other so as to form long fibers. If muscles move, the muscle fibers must slide relative to one another, and they do so by continuously forming protein-protein contacts between each other. One fiber pulls the other to exert the muscular force, then the protein-protein pairs dissociate again, in order to prepare for the next step. Similarly, almost everything that happens in the human body is driven by proteins that find and bind each other – to transmit information or build up living matter. “As yet, nobody has been able to observe how proteins associate in detail,” says Prof. Noé. The problem is that proteins are extremely small, about a billionth of a meter, and their structures fluctuate a lot, making this process invisible to microscopes, and it thus is not possible to take a "video" of how it happens.
The research groups of Prof. Frank Noé at Freie Universität Berlin and Dr. Gianni De Fabritiis at University Pompeu Fabra Barcelona have collaborated to produce what is the first atomic-detail computer simulation of protein-protein association and dissociation. According to the scientists, the main challenge was that atomic-detail molecular dynamics are incredibly expensive to simulate. The interactions of about 100,000 atoms need to be computed, and the force felt by every atom is evaluated. Then, the simulation moves every atom by a small distance in the direction of the force while it advances the simulated time by a femtosecond – a tiny fraction of a second. Each such step is computationally expensive, but a billion times a billion of such steps need to be performed in order to reach the typical duration that the proteins stay bound before they dissociate again. “The problem is that proteins are really sticky,” Prof. Frank Noé explains.
The world's fastest special-purpose supercomputer for molecular dynamics simulation, Anton 2, built by D. E. Shaw research, would require more than 10,000 years in order to simulate this process by brute force. “It is the type of high-risk project that is very difficult to get funding for, because when we started, no-one would believe that it's even possible,” says Noé.
Key to the success of the Berlin-Barcelona team was the combination of several new technologies that enabled a “divide and conquer” approach to the problem. GPUgrid, a distributed network developed by De Fabritiis' group was employed to collect compute time on graphics cards “donated” by volunteers around the globe. Thousands of short simulations were conducted that way, coordinated by a novel machine learning algorithm in such way that the overall protein association process could be simulated within one year instead of having to wait 10,000 years. Markov modeling, a method pioneered by Frank Noé and colleagues, was used to combine the many short simulations to an overall dynamical model that describes protein association and dissociation in full detail.
The project was supported by the European Research Council (ERC), the German Research Foundation (DFG), the Spanish Research Council (CSIC), and the Einstein Foundation Berlin.
Media Images
The photos may be downloaded by members of the media. They may be used free of charge when reporting in the context of the press release, provided that due credit is given the source.TOC Figure Caption
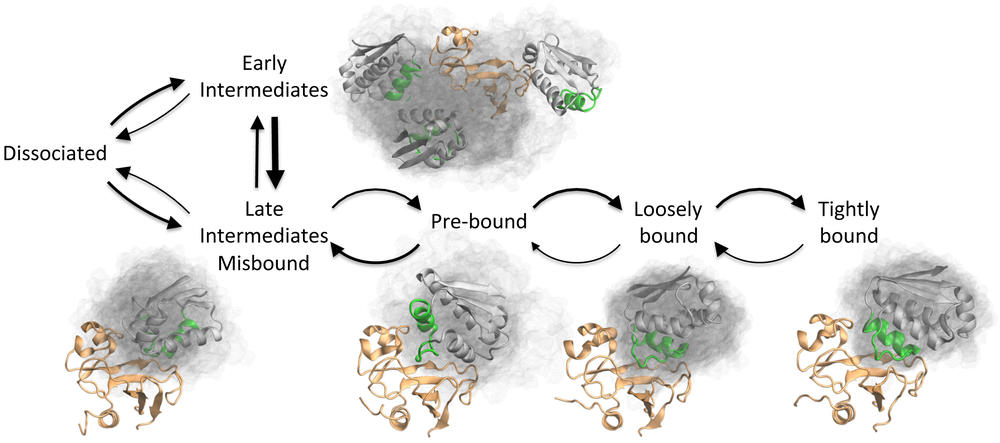
TOC Figure Caption: The association and dissociation process of the two bacterial proteins Barnase and Barstar has been reconstructed using unprecedented computer simulations. The image shows a few representative structures in which Barnase (orange) and Barstar (grey, transparent cloud indicates the flexibility of the protein in each state) can interact. The binding and dissociation process consists of a random jump process in this network of states, where the thickness of each arrow corresponds to the probability to transition along this direction.
Process of protein association and dissociation
Further Information
Publication
Nuria Plattner, Stefan Doerr, Gianni De Fabritiis, and Frank Noé (2017): “Complete protein–protein association kinetics in atomic detail revealed by molecular dynamics simulations and Markov modelling.” In: Nature Chemistry. DOI: 10.1038/NCHEM.2785
Contact
- Prof. Dr. Frank Noé, Department of Mathematics and Computer Science, Freie Universität Berlin, Tel.: +49 30 838 75354, Email: frank.noe@fu-berlin.de
- Dr. Nuria Plattner, Department of Mathematics and Computer Science, Freie Universität Berlin, Tel.: +49 30 838 75879, Email: nuria.plattner@fu-berlin.de